Plasmonic Nanopores
Biomolecules, like protein and DNA control all the processes that allow us to live. When they malfunction, you're in trouble and we better figure out what's going wrong. But that is not always particularly easy: from many biomolecules we do not understand how they work and why they work so well. To investigate that we should be to inspect them in close-up so that we can observe their function and intricate machinery that makes their little clocks go round.
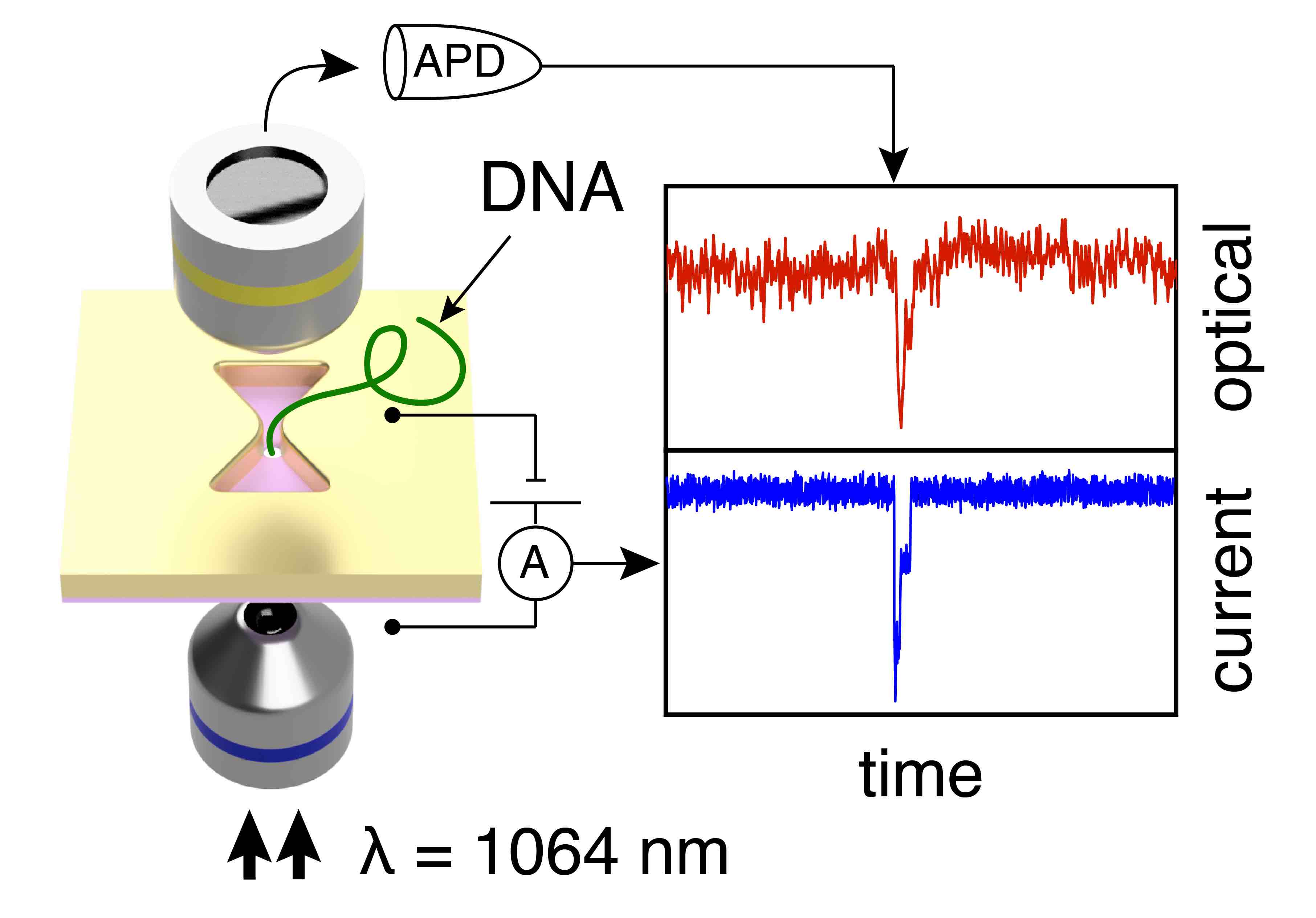
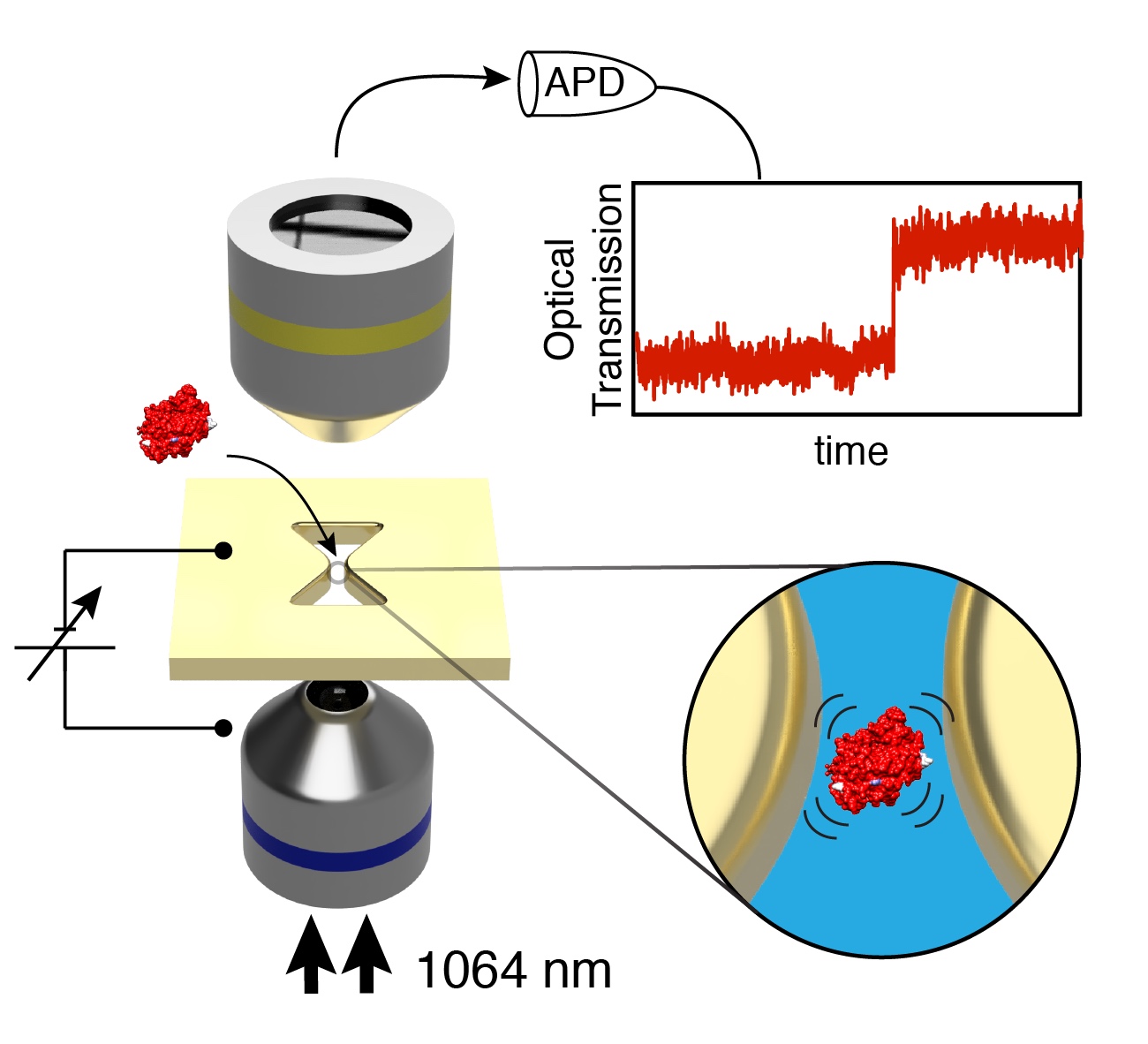
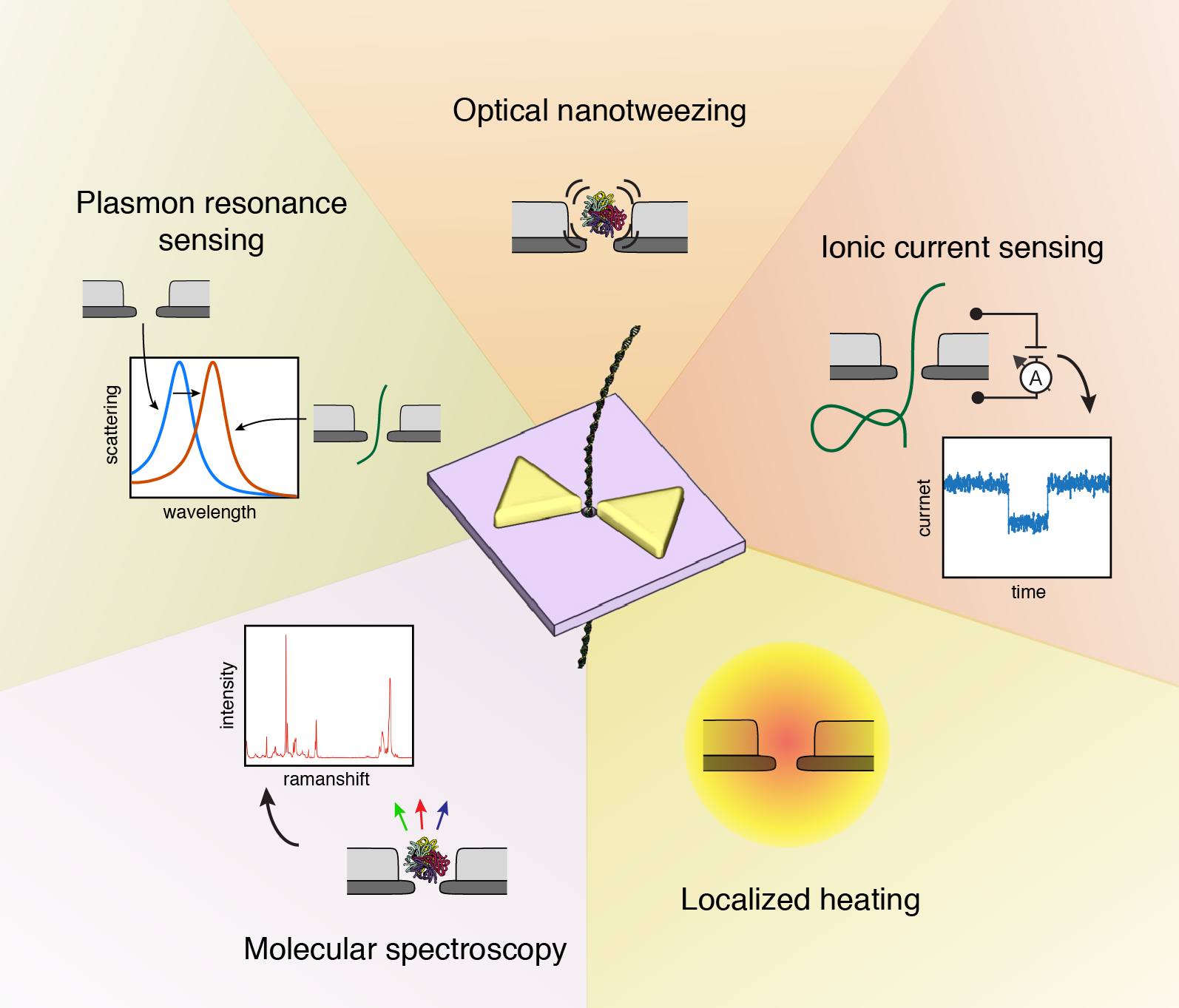
But grabbing a single molecule and inspecting its contents is really hard. Apart from the small size of the objects, biomolecules shake, shimmer, and bounce around a tremendous amount. How can one gently control something that small (without squashing or destroying it) and still be able to tell what it is? I have worked on a practicle solution to tackle that problem during my doctoral research: the plasmonic nanopore sensor to investigate and manipulate single biomolecules. The plasmonic nanopore is constructed from two single-molecule sensing devices merged into one: a solid-state nanopore, a tiny hole in a thin membrane that confines a static electric field, and a plasmonic nanoantenna, a gold nanostructure that concentrates light into nanoscale volumes (hotspots). Using these localized static and optical fields, biomolecules can be captured, trapped, perturbed, manipulated, and probed in a variety of ways. All controlled at will by the experimenter, one single molecule at the time.